Home
The introduction and overview of Fast Amber Rescoring for PPI inhibitors (farPPI) websever is available at “Home” page. In addition, the status of server is monitored on this page.

Job submission
A rescoring job can be submitted on the “Submit” page by clicking the “Submit” button when correct (1) Project name , (2) Docked pose file, (3) Receptor file, (4) Force field, and (5) Rescoring procedure are provided.

Job monitoring and result downloading
After a job is successfully submitted, the detailed information about it can be observed on the “Queue” page. When the calculation is done, a link for the corresponding result will be generated for downloading.

Datasets downloading
The ready-to-dock and ready-to-rescore benchmark datasets can be download on the “Download” page by clicking the corresponding links.

Tutorial for rescoring docking poses locally with MM/GB(PB)SA methods
In you need to rescoring docking poses locally which a workflow similar to farPPI, below is a concise and practical tutorial for you. Before you start this tutorial please download the material that will be used in this tutorial tutorial.zip.
Program version requirements: AmberTools/Amber 17 or higher, Python 2.7.14 or higher with matplotlib and seaborn libraries.
Step1: System preparation
Input: protein.pdb, poses.mol2
Output: complex_solv_radii.prmtop, complex_solv_radii.inpcrd
Script: step1_preparation.py with 5 position parameters pose_file receptor_file gaff|gaff2 ff99SB|ff14SB all|pb1|pb3|pb4|gb1|gb2|gb5|gb6
tar xvf tutorial.tar.gz
cd work/step1_preparation
python ../../script/step1_preparation.py ../../structure/poses.mol2 ../../structure/protein.pdb gaff2 ff14SB all
Step2: complex minimization
Input: complex_solv_radii.prmtop, complex_solv_radii.inpcrd
Output: min3.restrt
Script: step2_minimization.py with 2 position parameters preparation_dir all|pb1|pb3|pb4|gb1|gb2|gb5|gb6
cd ../step2_minimization
python ./../script/step2_minimization.py ../step1_preparation all
Step3: MM/GB(PB)SA rescoring
Input: complex.pdb, complex_solv_radii.prmtop, complex_radii.prmtop, receptor_radii.prmtop, ligand_radii.prmtop, min3.restrt
Output: mm_pbsa_statistics.out
Script: step3_rescoring.py with 3 position parameters preparation_dir receptor_file all|pb1|pb3|pb4|gb1|gb2|gb5|gb6
cd ../step3_rescoring
python ../../script/step3_rescoring.py ../step1_preparation ../../structure/protein.pdb all
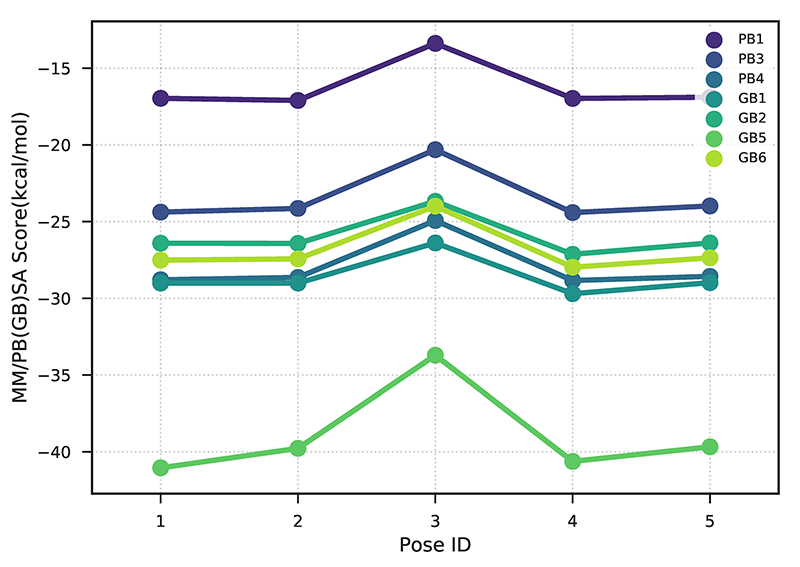
Step4: Data visualization
Input: mm_pbsa_statistics.out
Output: rescoring_result.csv, rescoring_result.pdf
Script: step4_visualization.py with 1 position parameter rescoring_dir
cd ../step4_visualization
python ../../script/step4_visualization.py ../step3_rescoring