What is NavDB?
NavDB (Navs Database) is an advanced, comprehensive database focused on sodium channel research,
offering extensive modulator and target information. It includes 8,023 data records, comprising 3,508
chemical modulators, 69 drugs, and 1,591 peptides, sourced from publicly available databases,
academic literature, and patents. All data have been rigorously curated to ensure accuracy and reliability.
Key Features of NavDB:
✔ Comprehensive Data: Provides a wide range of sodium channel modulator data, including small molecules, toxins, peptides,
and drugs, offering diverse information for researchers.
✔ Advanced AI Integration: Utilizes AI tools like AlphaFold and ADMETlab to enrich compound property data, enhancing the
depth and applicability of the database.
✔ Manual Curation of Binding Site Information: Through in-depth review of primary literature, NavDB includes detailed
annotations of compound binding sites (e.g., VSDII, VSDIV regions), a critical feature absent in other databases and highly valuable for structure-based drug design.
✔ Multi-Modal Search Capabilities: NavDB supports a range of advanced search methods, including text-based queries
(e.g., compound names, PubChem CIDs), structure-based search (e.g., input SMILES, draw molecules). These flexible tools enable researchers to
efficiently explore chemical, structural, and sequence-level data relevant to VGSC modulation.
✔ 3D Visualization: Features a 3D interactive interface, ensuring full visibility of sodium channel targets
and peptide structures, helping users intuitively understand and analyze the data.
✔ Easy Access: All data are freely available for download, supporting researchers in sodium channel studies,
drug development, and related fields.
NavDB is a vital resource for researchers and pharmaceutical companies in fields such as
pharmacology, drug discovery, and biology. It not only aids in advancing sodium channel research but also promotes the development of new therapeutic
strategies and fosters scientific collaboration and innovation worldwide.

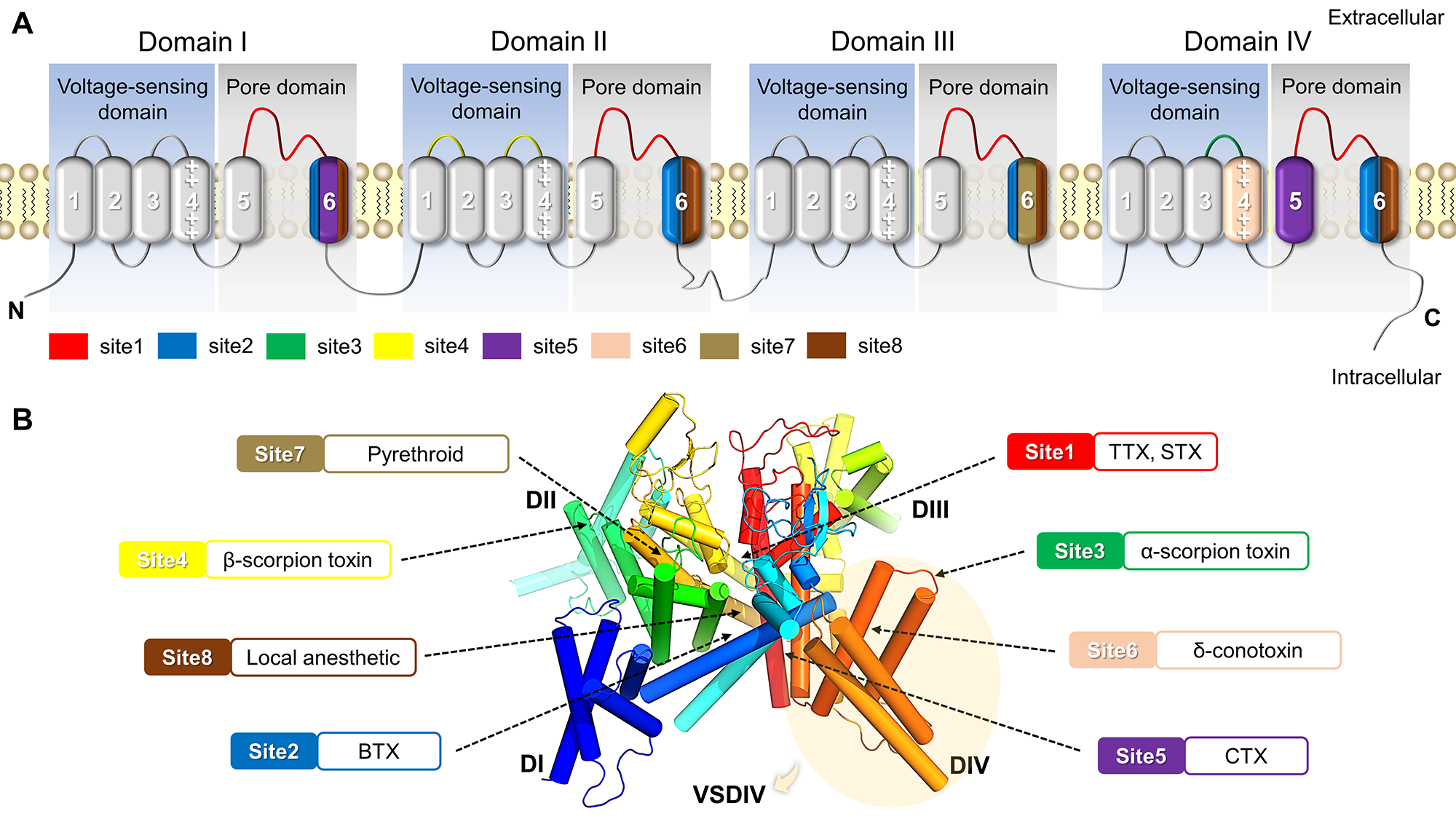
Binding Site
✔ Site1~8: Up to now, eight binding sites, which can interact with natural toxins and therapeutic drug molecules,
have been identified in the structures of VGSCs, and exhibit diverse physiological functions in response to different ligands.
✔ VSD: The VSD (voltage-sensing domain) is the important part of VGSCs, which can sense the change of external voltage and
adjust the opening of channel when the membrane is depolarized. The eukaryotic VGSCs have four VSDs (VSDI~VSDIV).
In recent years, the fourth VSD (VSDIV) has been determined as a new binding domain and several compounds targeting
this domain have entered clinical trials.
In NavDB, the binding site information of compounds are supplemented according to the above domain.
| Site | Ligand | Type | Binding domain |
|---|---|---|---|
| 1 | Tetrodotoxin (TTX), saxitoxin (STX), μ-contoxins | Extracellular pore blocker | DI~DIV P-loops |
| 2 | Batrachotoxin (BTX), veratridine (VTD), grayanotoxin (GTX), aconitine (ACT) | Intracellular pore gating activator (state dependent) | Near DI~DIV S6 (four repeats) |
| 3 | α-scorpion toxins, sea anemone toxins | Extracellular gating activator | Extracellular loops of DIV S3~S4 |
| 4 | β-conotoxin, δ-palutoxins, spider toxins | Extracellular gating blocker | Extracellular loops of DII S1~S2 and S3~S4 |
| 5 | Ciguatoxins (CTX), brevetoxins (PbTxs) | Intracellular pore gating activator (state dependent) | Near DI-S6 and DIV-S5 |
| 6 | δ-conotoxins | Extracellular gating activator | Near DIV-S4 |
| 7 | Pyrethroids | Intracellular gating activator | Near DIII-S6 |
| 8 | Small-molecule drugs, local anesthetics (LA) | LA binding site | Near DI~DIV S6 |
Biological Activity

Search
 (2) Advanced Search: NavDB provides a function for advanced searches. A large number of compounds can be searched by selecting
the molecular weight, SMILES length, organism, and (IC50). Users can download data in batches.
(2) Advanced Search: NavDB provides a function for advanced searches. A large number of compounds can be searched by selecting
the molecular weight, SMILES length, organism, and (IC50). Users can download data in batches.
 (3) Sequence similarity search: This feature is specifically designed for peptide searches. Users need to use this function under
the peptide page, where they can input a peptide sequence for matching. The results will be returned in a list format, sorted by
relevance from high to low.
(3) Sequence similarity search: This feature is specifically designed for peptide searches. Users need to use this function under
the peptide page, where they can input a peptide sequence for matching. The results will be returned in a list format, sorted by
relevance from high to low.

Target Page



Compound Page


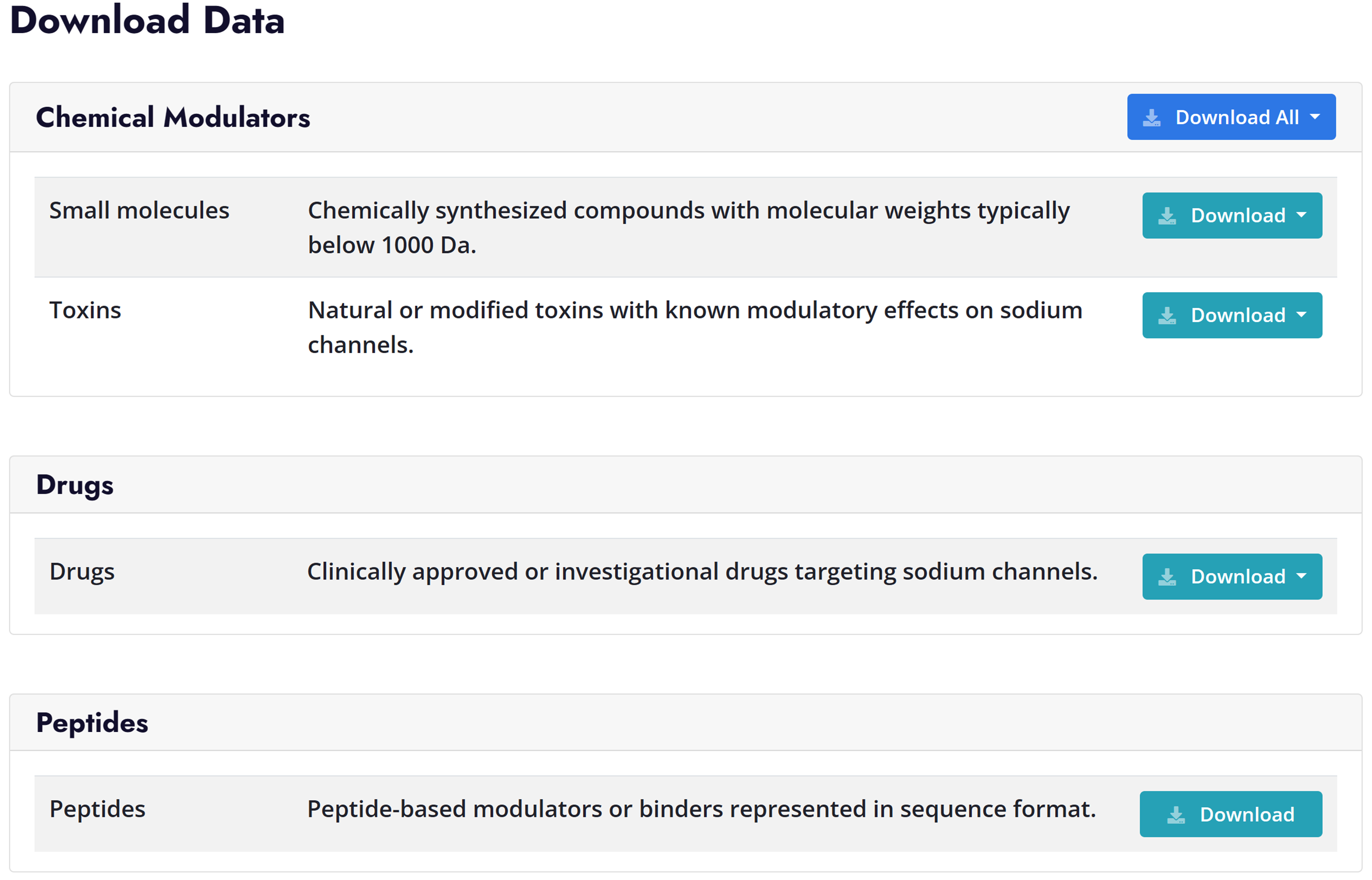
Peptide Page
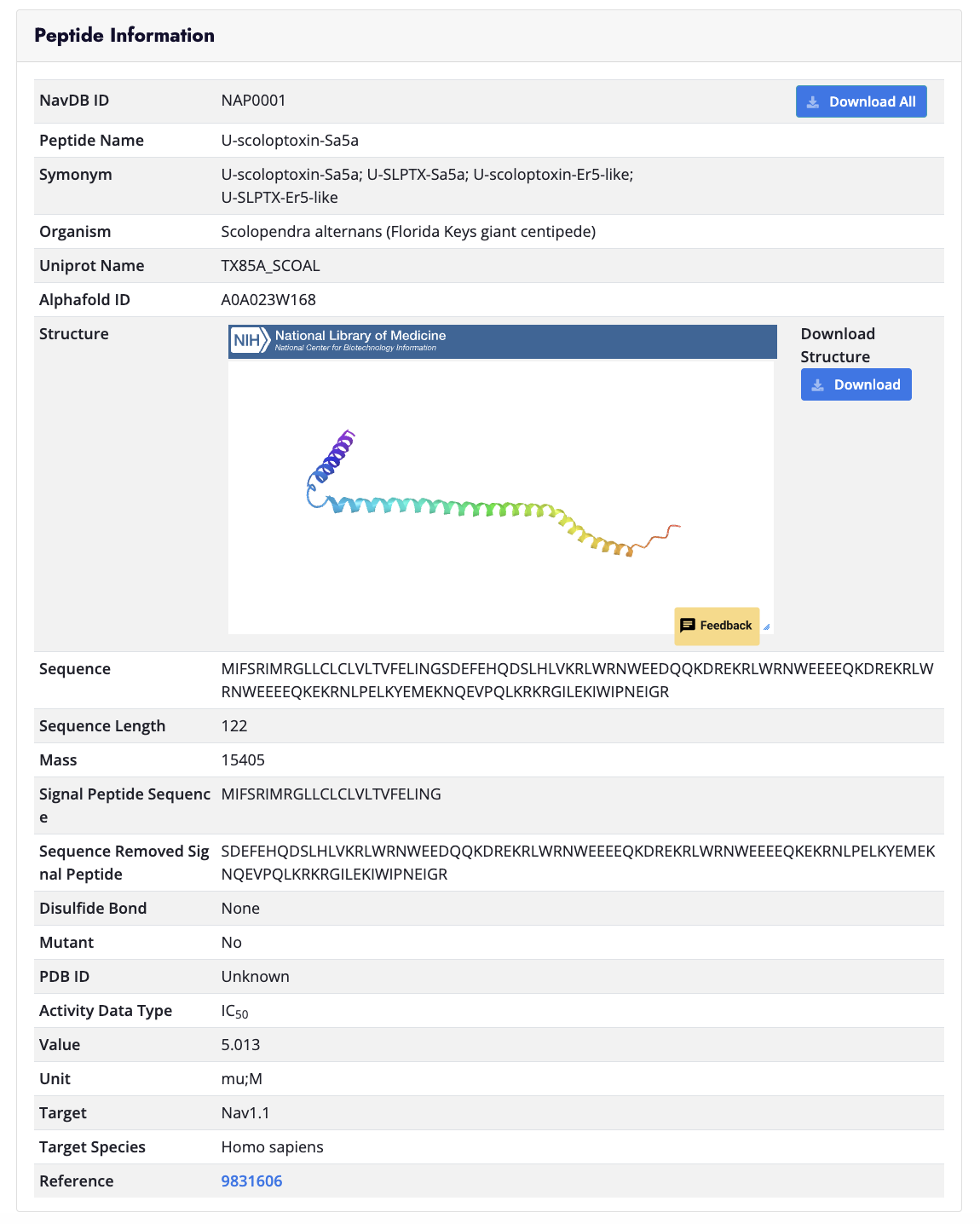
Drug Page
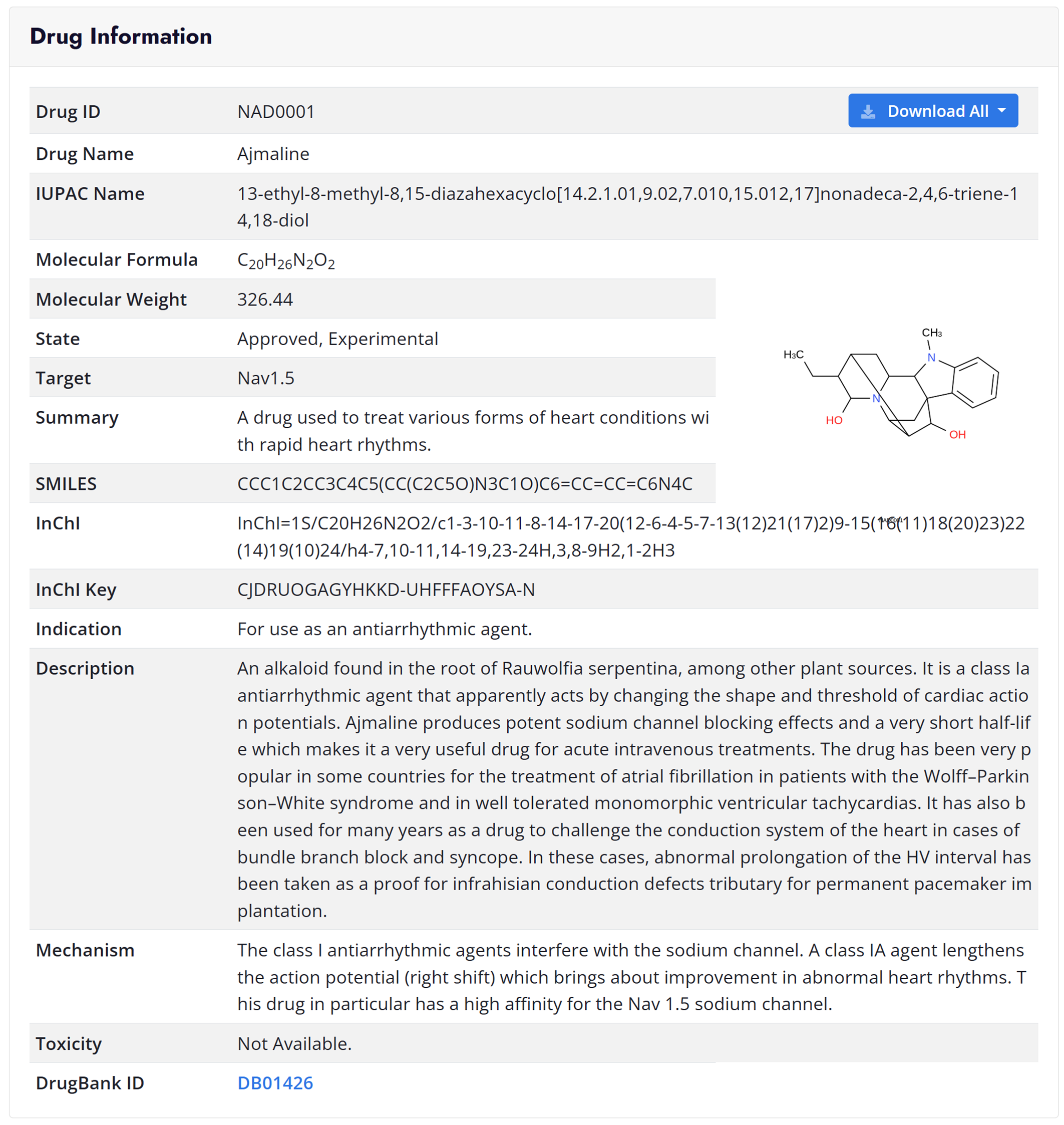
Download Page